Install the required packages and link to the correct github repository.
library(remotes)
remotes::install_github(repo="afitz-gmu/DGE-Analysis", build_opts = c("--no-resave-data", "--no-manual"))
library(GenomicVisualization)
Basic function to perfrom Differential Gene Expression by EdgeR, DESeq2 and Voom.
data("count.table")
design <- data.frame("trt" = colnames(count.table))
rownames(design) <- design$trt
design$trt <- as.integer(grepl("T",design[,1]))
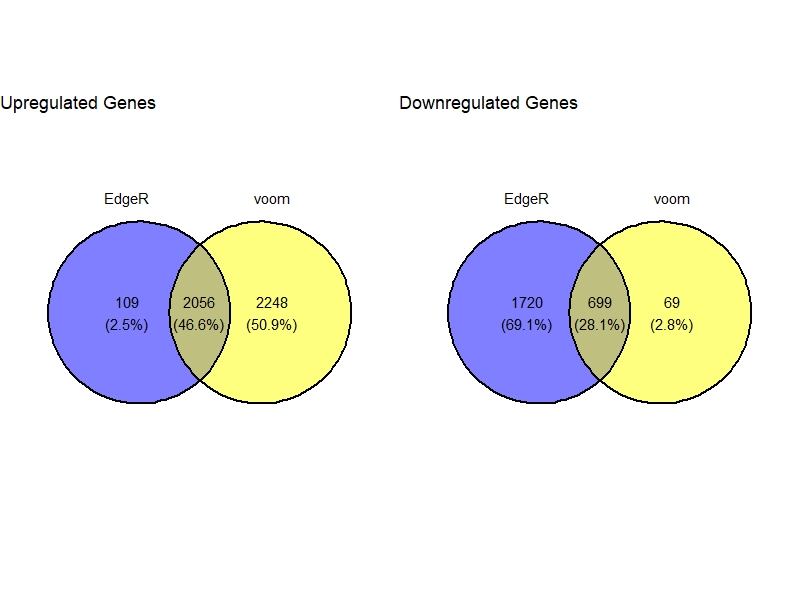
DGE.list <- DGE(count.table = count.table , design.matrix = design , method=c("EdgeR","voom") )
VennDig(DGE.list)
Prepare the data.
data("DEG")
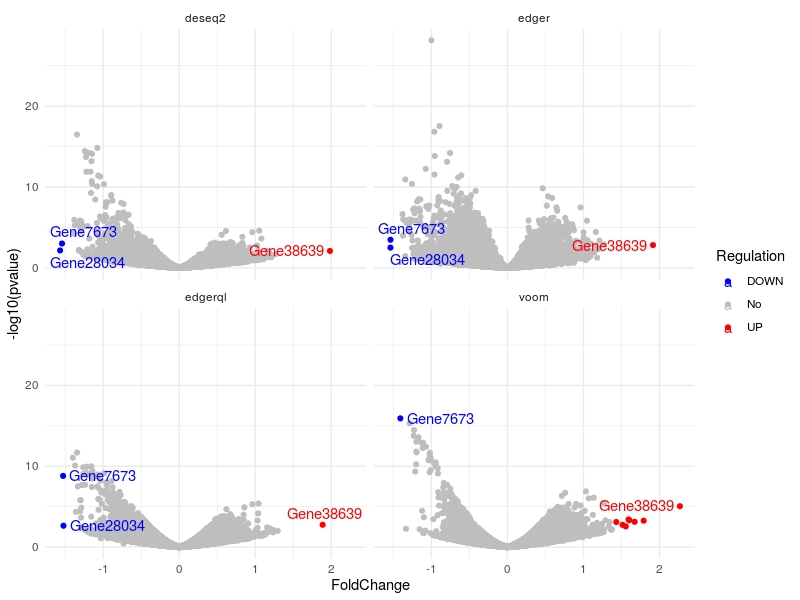
DE.list<-list("edger" =dge_edger, "edgerql" = dge_edgerql, "deseq2" = dge_deseq2, "voom" = dge_voom )
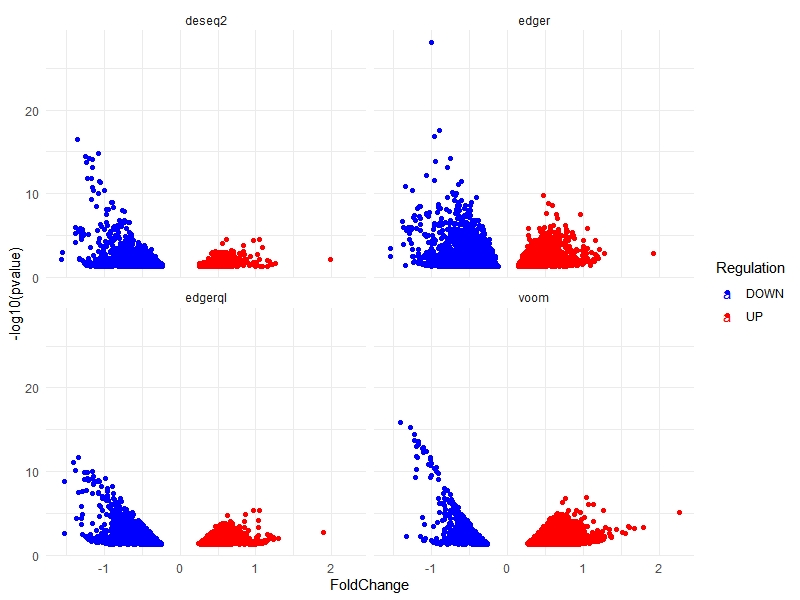
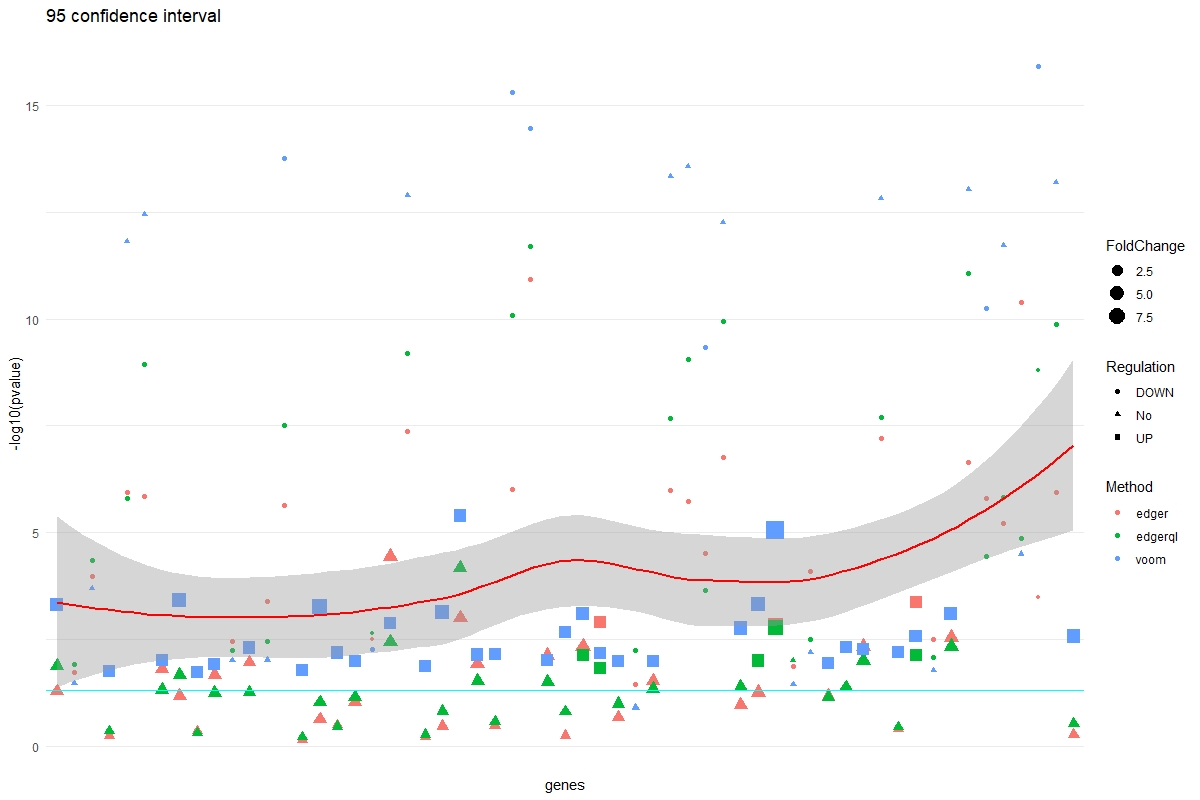
multiVolcano(DE.list, FoldChange = 1.4, DE.Only = FALSE , show.genes = TRUE )
For user defined gene list
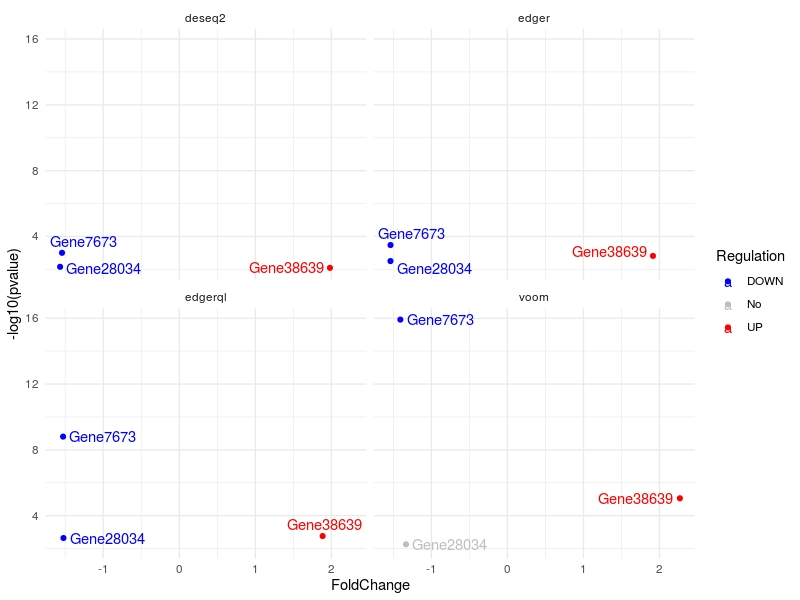
genes.list<-c("Gene7673", "Gene28034", "Gene38639")
multiVolcano(DE.list, FoldChange = 1.4, DE.Only = FALSE , genes = genes.list , show.genes = TRUE)
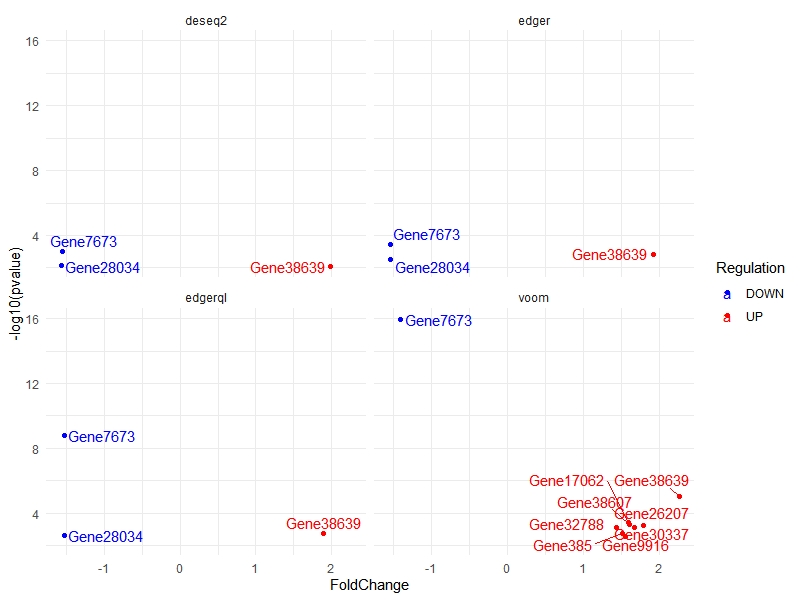
For showing Differentially expressed genes. This command will show only a few of the outlier genes)
multiVolcano(DE.list, show.genes = FALSE)
For showing Differentially expressed genes (marking show.genes as false for more asthetic plot)
multiVolcano(DE.list, FoldChange = 1.4 , show.genes = FALSE)
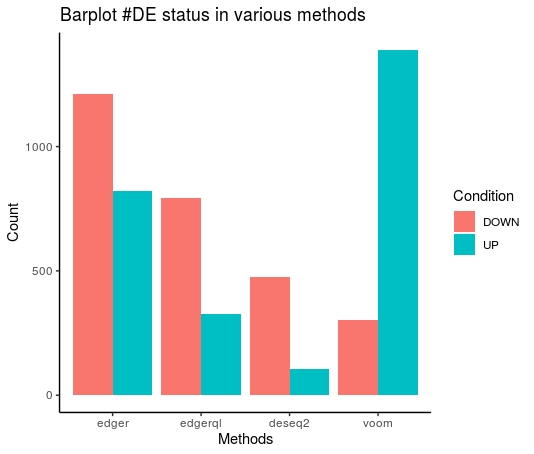
TabulateStats(DE=DE.list , FoldChange=0 , cutoff =0.01 )
| edger | edgerql | voom | |
|---|---|---|---|
| DOWN | 1210 | 794 | 302 |
| No | 33947 | 34858 | 34287 |
| UP | 821 | 326 | 1389 |
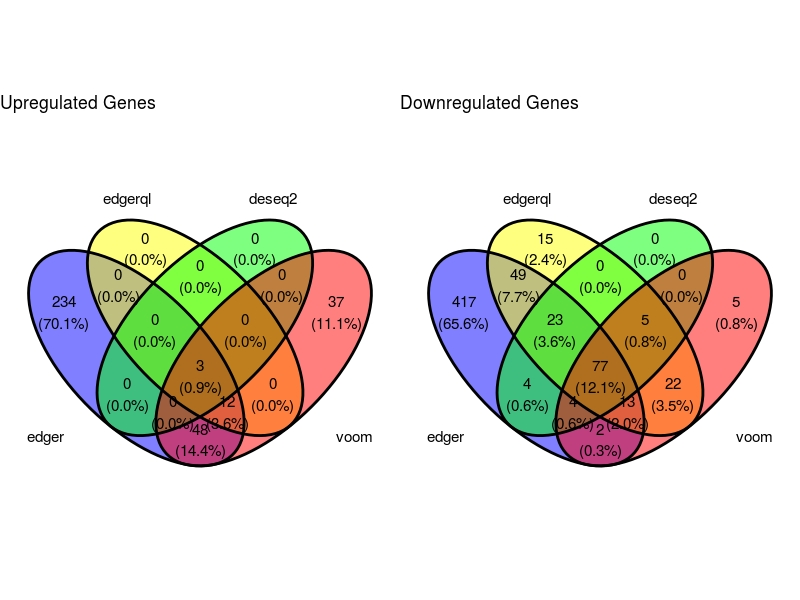
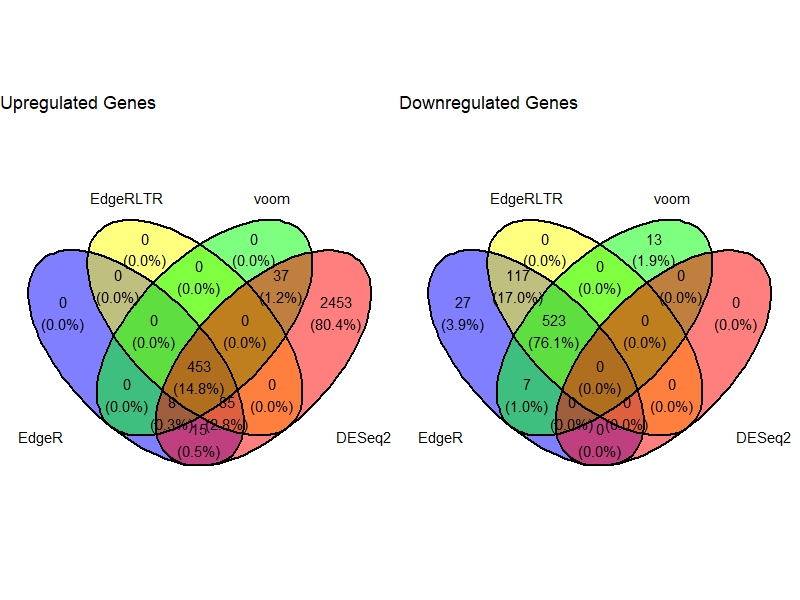
VennDig(DE=DE.list , FoldChange=1.5 , cutoff =0.01 , type.sig ="FDR")
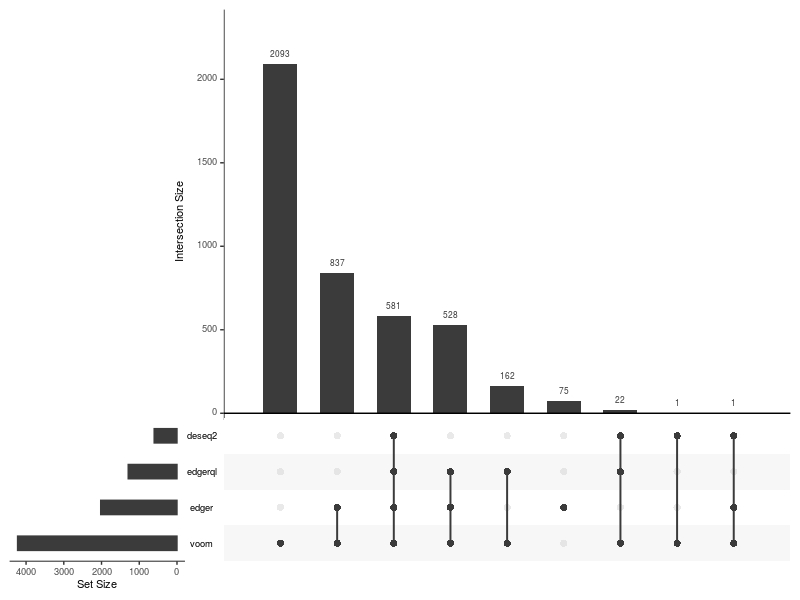
UpSetPlot(DE=DE.list , FoldChange=0 , cutoff =0.01 , type.sig = "p")
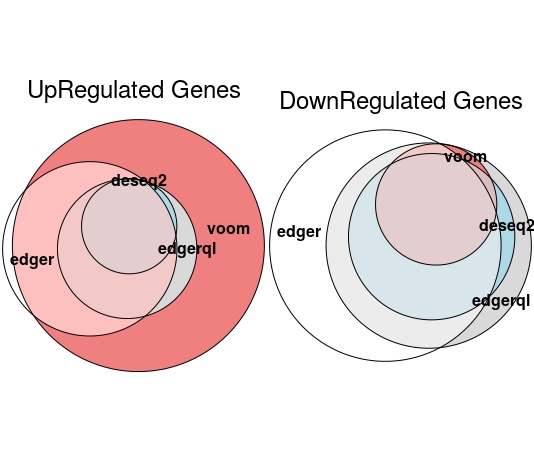
EulerrPlot(DE=DE.list , FoldChange=0 , cutoff =0.05 , type.sig ="p")
DGE.CI(DE=DE.list , FoldChange=1.2 , cutoff =0.05 , type.sig ="p")
| edger FoldChange | edgerql FoldChange | voom FoldChange | Min | Max | edger pvalue | edgerql pvalue | voom pvalue | Min | Max | |
|---|---|---|---|---|---|---|---|---|---|---|
| Gene11876 | 0.253268010646449 | 0.25665960045484 | 0.333519105983995 | 0.253268010646449 | 0.333519105983995 | 0.00966910768271969 | 0.0138679244791296 | 0.0390346542589136 | 0.00966910768271969 | 0.0390346542589136 |
| Gene15604 | 0.27180343736292 | 0.271787631013647 | 0.302097146830169 | 0.271787631013647 | 0.302097146830169 | 0.000348901761561591 | 0.00103259777034587 | 3.97342702281636e-09 | 3.97342702281636e-09 | 0.00103259777034587 |
| Gene15888 | 0.30094351744471 | 0.300302928429722 | 0.332617475249204 | 0.300302928429722 | 0.332617475249204 | 0.000405266030230859 | 2.99397057448929e-06 | 1.16326308848636e-09 | 1.16326308848636e-09 | 0.000405266030230859 |
| Gene22288 | 0.263474804206977 | 0.263400660850057 | 0.2916799978793 | 0.263400660850057 | 0.2916799978793 | 0.0005861510066759 | 4.17234374228856e-05 | 1.5530006115196e-10 | 1.5530006115196e-10 | 0.0005861510066759 |
| Gene29009 | 0.294253032199853 | 0.293993599013282 | 0.330000812105133 | 0.293993599013282 | 0.330000812105133 | 2.62505647540443e-05 | 2.81215059098373e-06 | 5.0579118548581e-10 | 5.0579118548581e-10 | 2.62505647540443e-05 |
| Gene30812 | 3.0011502354894 | 2.82286115292464 | 3.54654652335402 | 2.82286115292464 | 3.54654652335402 | 0.0443053766662665 | 0.0192060783610423 | 0.00229564569695183 | 0.00229564569695183 | 0.0443053766662665 |
| Gene32136 | 0.253951391140971 | 0.253847733944933 | 0.276660037325252 | 0.253847733944933 | 0.276660037325252 | 0.000311904095164778 | 8.10948071015179e-07 | 8.89107509409887e-12 | 8.89107509409887e-12 | 0.000311904095164778 |
| Gene3219 | 0.26132542426818 | 0.260852100787794 | 0.292821559694857 | 0.260852100787794 | 0.292821559694857 | 3.9108014179455e-08 | 7.25440228196772e-08 | 4.04635632813972e-11 | 4.04635632813972e-11 | 7.25440228196772e-08 |
| Gene3636 | 0.276773935358202 | 0.276658009021903 | 0.30742464752486 | 0.276658009021903 | 0.30742464752486 | 0.000320786232115754 | 3.12890638531982e-05 | 2.80586403681253e-10 | 2.80586403681253e-10 | 0.000320786232115754 |
| Gene36720 | 0.282886374057711 | 0.283071871234626 | 0.310527439322474 | 0.282886374057711 | 0.310527439322474 | 0.000489633506875775 | 2.9117905222642e-06 | 1.90836815806636e-10 | 1.90836815806636e-10 | 0.000489633506875775 |
| Gene37809 | 0.274775812479113 | 0.274781151367946 | 0.297749013272049 | 0.274775812479113 | 0.297749013272049 | 0.00409971202637718 | 0.0403344717227362 | 5.83264594328474e-07 | 5.83264594328474e-07 | 0.0403344717227362 |
| Gene38199 | 0.295370543964787 | 0.296061444016233 | 0.328422674982562 | 0.295370543964787 | 0.328422674982562 | 8.2482427048141e-05 | 8.10948071015179e-07 | 1.52215838845913e-09 | 1.52215838845913e-09 | 8.2482427048141e-05 |
| Gene38639 | 6.7835229404579 | 6.58741660386054 | 9.6638046407115 | 6.58741660386054 | 9.6638046407115 | 0.0573934369490823 | 0.136024039700254 | 0.00415995911576501 | 0.00415995911576501 | 0.136024039700254 |
| Gene38937 | 0.283452115852866 | 0.27553856211236 | 0.315105821098576 | 0.27553856211236 | 0.315105821098576 | 0.00825053138204574 | 0.195476227936559 | 0.181147657675811 | 0.00825053138204574 | 0.195476227936559 |
| Gene40770 | 0.289698705765732 | 0.289780204987156 | 0.325317239975445 | 0.289698705765732 | 0.325317239975445 | 3.4623122963608e-05 | 3.08747716025276e-05 | 5.52436063074383e-10 | 5.52436063074383e-10 | 3.4623122963608e-05 |
| Gene6678 | 0.251819899040539 | 0.246647238865664 | 0.301947985426532 | 0.246647238865664 | 0.301947985426532 | 0.00010352524415427 | 1.56612785739337e-07 | 4.16666249706773e-10 | 4.16666249706773e-10 | 0.00010352524415427 |
| Gene6710 | 0.271459164291596 | 0.27145692542583 | 0.301120050678203 | 0.27145692542583 | 0.301120050678203 | 0.000430650590640714 | 0.0118339982149324 | 9.04425211253826e-08 | 9.04425211253826e-08 | 0.0118339982149324 |
| Gene720 | 0.273484715998214 | 0.273434784275442 | 0.302884849886368 | 0.273434784275442 | 0.302884849886368 | 0.00122751324077398 | 0.00102621047894686 | 4.53220294233197e-09 | 4.53220294233197e-09 | 0.00122751324077398 |
| Gene7339 | 0.284743280003845 | 0.275612814902474 | 0.32548462536702 | 0.275612814902474 | 0.32548462536702 | 1.22959638267496e-07 | 0.00588064870290072 | 0.0108920996649206 | 1.22959638267496e-07 | 0.0108920996649206 |
| Gene7673 | 0.21499523132271 | 0.216980068912695 | 0.244685085090055 | 0.21499523132271 | 0.244685085090055 | 0.0218327855690298 | 3.60045900816876e-06 | 4.35821145336272e-12 | 4.35821145336272e-12 | 0.0218327855690298 |
| Gene9230 | 0.282854723987658 | 0.282620482988113 | 0.310124743179966 | 0.282620482988113 | 0.310124743179966 | 0.000351248577561045 | 8.10948071015179e-07 | 3.29044695217997e-10 | 3.29044695217997e-10 | 0.000351248577561045 |
# Install the required packages
BiocManager::install("multtest")
library(multtest)
# load the golub dataset
data(golub)
ds<-data.frame("trt" = golub.cl)
colnames(golub) <-paste0("Name",1:length(golub.cl))
row.names(ds) <- colnames(golub)
ds$trt <- paste0("C",ds$trt)
xx<-as.data.frame(round(exp(golub)*1000))
DGE.list <- DGE(count.table = xx , design.matrix = ds )
VennDig(DGE.list)
DGE.CI(DE=DE.list , FoldChange=1.2 , cutoff =0.05 , type.sig ="p")