WIP @jsteb january 2024
At its core, the MRTWin framework and pipeline can be used to perform integrated MR sequence development and simulation of the MR images. The neat part of the pipeline is given by the ability to directly read in any valid .seq file and simulate the forward process.
The sequence development part is handled via Pypulseq (GitHub Repository). The simulation part is handled via the MRZeroCore package (GitHub Repository), where the ominous 👻 phase distribution graph (PDG) formalism is used. Peruse Magnetic Resonance in Medicine Preview site for the soon-to-be-released paper by Moritz Zaiss et al.
The backend makes use of the tensor machinery and automatic differentiation engine of PyTorch. The reconstruction of the data utilizes the torchkbnufft package (GitHub Repository) to perform non-uniform Fast Fourier Transform with Kaiser-Bessel gridding in PyTorch.
For ease of usage, we installed the MRTwin pipeline on the julia cluster machines. First gain access to the desired machine by following the tutorial written by Max. The first step is to join the JMU VPN network. You cannot access the julia cluster machines from the outside internet. Afterwards, basal access is usually achieved via ssh using the floating IP of the desired machine.
| Host Name | Floating IP | SSH Key Name |
|---|---|---|
| mrf1 | 10.106.246.104 | mrf1_key.pem |
| mrf2 | unknown | unknown |
⚠️ Warning Currently, we all operate as the same userubuntuon the variousjuliacluster machines. Make sure to not accidentally delete or corrupt any data of your colleagues 😱. Further, we have to coordinate the resources of the machines. It is a good idea to check that the machine you intend to use for your calculations is not currently occupied by a fellow researcher. Otherwise, resource fighting of the jobs/processes may dramatically slow down the computations. 🐌 You can check the current resource consumption in the terminal with thehtopcommand. If either CPU usage or memory consumption is high, you may have to select another host 💻 or contact the colleague to stop being a resource hog 🐗.
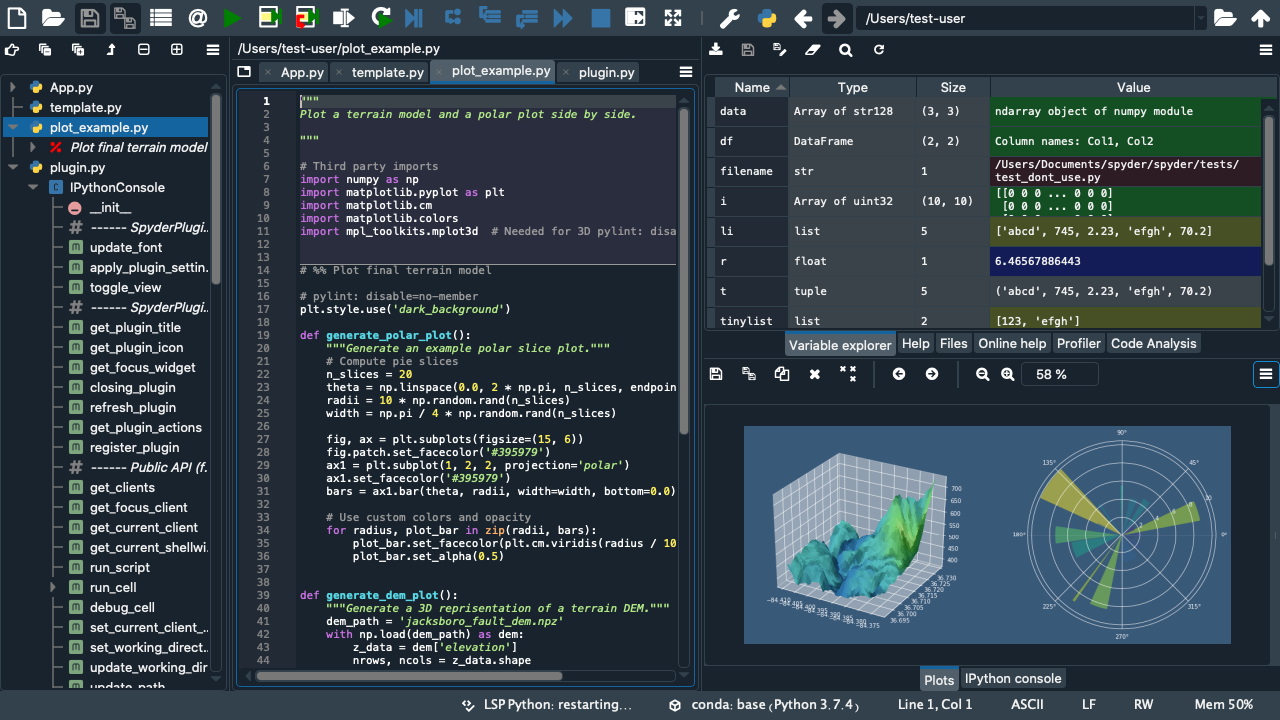
You can use the virtual network computing (VNC) protocol together with a suitable client (e.g. TigerVNC) to gain GUI access to one of the host machines. Then open a terminal session inside the Ubuntu GUI. Then you You should see something like the image below.
Use the conda environment activation tutorial to activate the mrtwin environment. Then you may use your desired development tool of choice, e.g. Spyder IDE (command spyder), Jupyter Notebooks (command jupyter notebook or jupyter nbclassic), VS Code (command code), etc. pp. to implement and simulate the new hottest MR sequence! 🔥
You can avoid the VNC protocol and sending GUI screen content over the network and setup a complete remote development environment! 🥳 This may provide a leaner setup and may be more responsive as well.
Depending on the desired development tools, you will set up the remote connection differently:
See the Spyder docs to connect to a remote kernel running on the julia cluster.
Tutorial: Link
The advantages over full GUI transmission are similar to the ones specified in jupyter notebook setup.
Connect to the julia cluster VM via the VS Code remote module. This allows both classic file-based development and Jupyter notebook-based development inside VS Code.
Tutorial: Link
Upon gaining shell access to one of the julia cluster machines, you can start a headless Jupyter notebook server with the commands:
jupyter notebook --no-browser --port=$YOURPORTfor the most recent Jupyterlab notebook experience or
jupyter nbclassic --no-browser --port=$YOURPORTfor the classic Jupyter notebook experience. The server is headless due to the --no-browser flag and expects connections via a user-specified port $YOURPORT (i.e. a free port, specified as an integer between 0 and 65535). The browser-based notebook experience allows the usage of cool and interactive tools like ipywidgets to live-modify sequence and simulation parameters! To access the notebook in your local browser, you have to create a ssh tunnel that correctly connects your local machine to the remote cluster machine where the jupyter notebook server runs.
When you want to access the jupyter notebook in your local browser (i.e. the hacker way ;)) then you need to create a ssh tunnel such that the network traffic is forwarded to the jupyter server on the remote julia cluster machine. The command to start the ssh tunnel is
ssh -N -f -L $LOCALPORT:localhost:$YOURPORT:localhost username@hostnamewhere the $LOCALPORT is the one you want to access on your local machine (it is advisable to use $LOCALPORT == $YOURPORT for ease of use) and $YOURPORT is the chosen port the jupyter server is listening to. Use the login credentials for the mrf cluster machine that were given to you by Max or Jannik. The arguments do the following: -N makes that no remote command is executed since the connection is established only for port forwarding, -f ensures that the tunnel is run continuously in the background and -L specifies that the given port on the local (client) host is to be forwarded to the given host and port on the remote side. Many of these steps can be automated via the ~/.ssh/config file such that you do not have to type them again or recall them tediously via your shell history. See e.g. the ssh manpages. If everything is working, you can access the jupyter notebook environment in your local browser under the address localhost:$LOCALPORT.
The julia cluster machines should have conda preinstalled. If not the case, contact Jannik for bugfixing 🐛🧰 purposes. In the bash terminal, you should see the conda environment prefix, in the above example (base), your user name (here ubuntu) and the current host name (here mrf1). Before starting off, you have to activate the correct conda environment to be able to access all package elements of the MRTwin. For this, you should activate the preinstalled environment mrtwin with the command
conda activate mrtwinA list of all conda environments can be printed via the command conda env list. The successful activation of the conda environment is reflected in the changed environment prefix in front of the user name and host name specification.
If you still have problems or open questions, ask Jannik for tech support