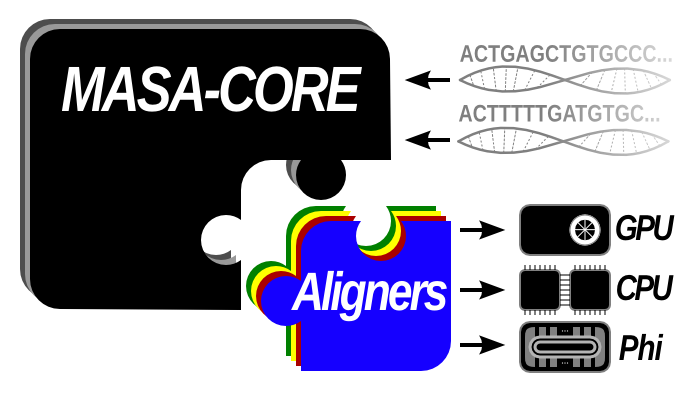
Multi-Platform Architecture for Sequence Aligner (MASA) is a flexible software framework that simplifies the creation of DNA sequence alignment applications in multiple hardware/software platforms. This framework supports the alignment of huge DNA sequences with more than 200 million base pairs (MBP).
The MASA-Core library is based on CUDAlign, a tool able to align huge sequences in CUDA capable GPUs. The MASA-Core contains 90% of the original CUDAlign source code, isolating the platform-independent features. Thus, any developer may create "aligners" for different hardware/software platforms, reimplementing only some methods of the original code and linking it against the portable MASA-Core library. Furthermore, new platform-independent optimizations may be deployed into the MASA-Core, leading to a wider contribution to all the specific aligners implementations. Platform-specific aligners will be maintained in separate repositories, with a static copy of the masa-core source.
The MASA-Core contains the following features:
- Produces optimal alignments for DNA sequences of any length (even greater than 200 MBP)
- Based on Smith-Waterman and Needleman-Wunsch algorithms
- Uses Myers-Miller strategy during the matrix computation (linear memory complexity)IPDPS2011
- Produces Local, Semi-Global and Global optimal alignments
- Multi-node support for homogeneousCCGRID2014 and heterogeneous hardwarePPOPP2014
- Block Pruning optimizationTPDS2012
- Portable C/C++ code. Compiled in Linux and MacOS X, for 32 and 64 bit platforms
Latest Version: masa-core-1.3.9.1024.tar.gz
tar -xvzf masa-core-1.3.9.1024.tar.gz
cd masa-core-1.3.9.1024
./configure
make
To create MASA executable files, download one of the extensions listed below. Each extensions project contains a copy of the MASA-Core source.
Each project that supplies a different Aligner linked with MASA-Core is called a MASA-Extension. Here is a list with some MASA-Extensions for some supported platforms.
- MASA-CUDAlign: Extension that uses CUDA NVidia GPUs (based on the original CUDAlign code)
- MASA-OpenMP: Uses OpenMP to computes many blocks in parallel using CPU or Intel Phi cores
- MASA-OmpSs: Uses OmpSs to run parallel tasks respecting dependencies
- MASA-Serial: This is a simple extension that runs with serial CPU code
./masa-extension [options] seq1.fasta seq2.fasta
All the command line arguments can be retrieved using the --help parameter. See the wiki for a list of command line examples.
See the doxygen pages.
MASA-Core is an open source project with public license (GPLv3). A copy of the LICENSE is maintained in this repository.
| [TOPC2016] | MASA: a Multi-Platform Architecture for Sequence Aligners with Block Pruning. TOPC:2(4):28. Edans Sandes, Guillermo Miranda, Xavier Martorell, Eduard Ayguadé, George Teodoro, Alba Melo. |
| [TPDS2016] | Genome Wide Alignment in GPU Cluster with Incremental Speculative Traceback. TPDS 2016. Edans Sandes, Guillermo Miranda, Alba Melo, Xavier Martorell, Eduard Ayguadé. |
| [CCGRID2014] | CUDAlign 3.0: Parallel Biological Sequence Comparison in Large GPU Clusters. CCGRID 2014:160-169. Edans Sandes, Guillermo Miranda, Alba Melo, Xavier Martorell, Eduard Ayguadé. |
| [PPOPP2014] | Fine-grain parallel megabase sequence comparison with multiple heterogeneous GPUs. PPOPP 2014:383-384. Edans Sandes, Guillermo Miranda, Alba Melo, Xavier Martorell, Eduard Ayguadé |
| [TPDS2013] | Retrieving Smith-Waterman Alignments with Optimizations for Megabase Biological Sequences using GPU. TPDS:24:5:1009-1021. Edans Sandes, Alba Melo |
| [IPDPS2011] | Smith-Waterman Alignment of Huge Sequences with GPU in Linear Space. IPDPS 2011: 1199-1211. Edans Sandes, Alba Melo |
| [PPOPP2010] | CUDAlign: using GPU to accelerate the comparison of megabase genomic sequences. PPOPP 2010: 137-146. Edans Sandes, Alba Melo |