- Data from DNA or RNA.
- 4 letters: A, C, G, T
- Data from Proteins.
- IUPAC: The 20 standard amino acids: A, R, N, D, C, Q, E, G, H, I, L, K, M, F, P, S, T, W, Y, V.
- Extended IUPAC: 26 letters. In addition to the standard 20 amino acids, this includes: B, X, Z, J, U, O.
- X is for Unknown or "other" amino acid.
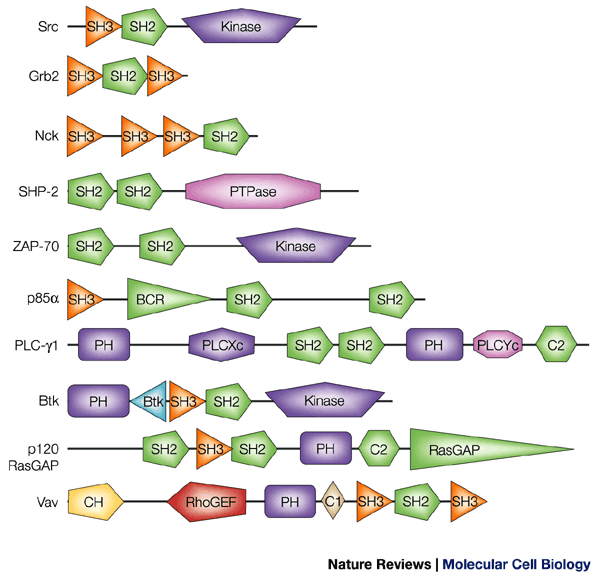
- Domains are sequences that are evolutionarily conserved, and as such have a well-defined fold and function.
- Pfam is a database of 32,207,059 protein domains used extensively in bioinformatics.
- Sequences in Pfam are clustered into evolutionarily-related groups called families.
Read about Ramachandran plot
- GO
- Uniprot
Grafo de interacción de proteinas.
Basic Local Alignment Search Tool. Algoritmo que determina el nivel de similaridad de 2 secuencias. Puede usarse:
- A nivel de secuencias de nucleótidos (Nucleutide BLAST)
- A nivel de secuencias de aminoácidos (Protein BLAST)
- UniProt: Secuencias de aminoácidos.
- UniProtKB: UniProt Knowledgebase
- Swiss-Prot: Manually annotated and reviewed (561.911)
- TrEMBL: Automatically annotated and not reviewed. (177.754.527)
- UniRef: UniProt Reference Clusters
- UniRef50: 50% (39,178,216)
- UniRef90: 90% (107,153,647)
- UniRef100: 100% (216,491,817)
- UniParc: UniProt Archive
- UniProtKB: UniProt Knowledgebase
- Pfam: Secuencias aminoácidos + dominios.
- Protein data bank: Muy pocas con respecto a UniProt. A parte de la secuencia contiene metadatos y repr. 3D.
- En kaggle: 400000 protein data set (146 MB)
Unsupervised learning is a pre-task that allows the neural net to learn from lot of unlabeled data.
- Language Model (LM): Predict next aminoacid. (used in RNNs)
- Masked Language Model (MLM): Predict hidden aminoacids. (introduced in BERT paper)
- Replaced Token Detection: Is this aminoacid real or fake? (introduced in ELECTRA)
- Secondary structure prediction
- Contact prediction: Predic in a pair of aminocis are within 8 angstroms of each other.
- Remote homology detection.
- Count vectorizers with n-grams
- TF-IDF
- Ultra-Deep Learning Model (2016)
- AlphaFold: From seq -> predict 3D shape
- Paper in Nature (Jan 2020)
- Paper in Proteins (Sep 2019)
- DeepDom (January 2019)
LSTM - UniRep (March 2019)
mLSTMUnsupervised - UDSMProt (September 2019)
AWD-LSTMUnsupervised - Kaggle: LSTM with keras
- Character-Level LSTM in PyTorch
- Protein-Structure-Prediction (weights and biases)
- Read Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences: After training the Transformer algorithm to process amino acid sequences, researchers looked at the embedding learned by the model. Above all, they found out that the neural network had built a complex representation of the input sequences. This in turn reflects their biological properties such as activity, stability, structure, binding etc. In other words, the deep learning algorithm learned important biochemical properties characterising the different amino acids and proteins, all by itself, without any supervision.
- Illustrating the Reformer
- ELECTRA: Otra forma de hacer el aprendizaje no supervisado
- Advices for training transformers: Train Large, Then Compress twitt, blog, paper
- Kaggle winner solution to Google’s QUEST Q&A Labeling: BERT, RoBERTa, BART
- Andrés Solution to Predicting Molecular Properties
- Choose one from Hugging face
- Train with fastai
- 3D protein