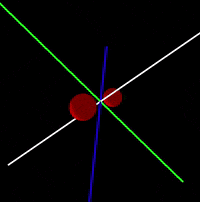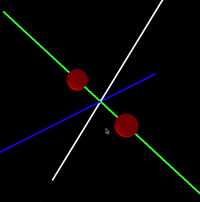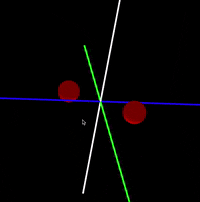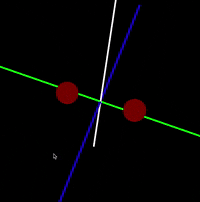
Java/Processing 3 application that simulates interaction between atoms in a molecule
The classes shown below are abstracted and only shows the most important variables and methods
class Nucleus {
int AtomicNumber;
String symbol, name;
double AtomicWeight, mass;
int numberOfProtons, numberOfElectrons;
double radius;
double electronegativity, firstIonizationEnergy;
int[] oxidationStates;
int[] RGB;
}This class handles initializing the properties of each atom object
class Electron {
int n, l, m, s;
void returnQuantumNumbers();
}This class handles initializing the properties of each electron ojbect and can return the values
class Atom extends Nucleus {
HashMap<String, List<Electron>> Quantum;
int charge;
int numberOfElectrons;
int energyLevels;
int x, y, z;
void setUpOrbitals();
void returnOrbital();
String[] returnProperties();
int getValanceElectrons();
}This class joins the nucleus and electron classes together to make an Atom object. The class also holds the x, y, z positions of the atom for when modelling them in 3-Dimensional space
class Bond {
Atom atom1, atom2;
int numberOfBonds;
void addBond();
double getBondPolarity();
}This class is responsible for creating the bond information between two atoms
class Molecule {
ArrayList<Atom> atoms;
String[] atomNames;
void dissectMolecule(String molecule);
int findAtomicNumber(String AtomicSymbol);
public String[] getAtoms();
}This class is responsible for dissecting the user's input molecule and breaking each character into a separate atom object with the method dissectMolecule(String molecule). With each atomic symbol broken down, the atomic number can be found with findAtomicNumber(String AtomicSymbol).
class Startup {
void startupWindow();
}This class contains all of the methods that displays the menus and grabs user input. Other windows within this class displays information about the molecule entered
class Environment extends PApplet {
void setup();
void draw();
}This class uses the library processing and peasy cam to 3-D model