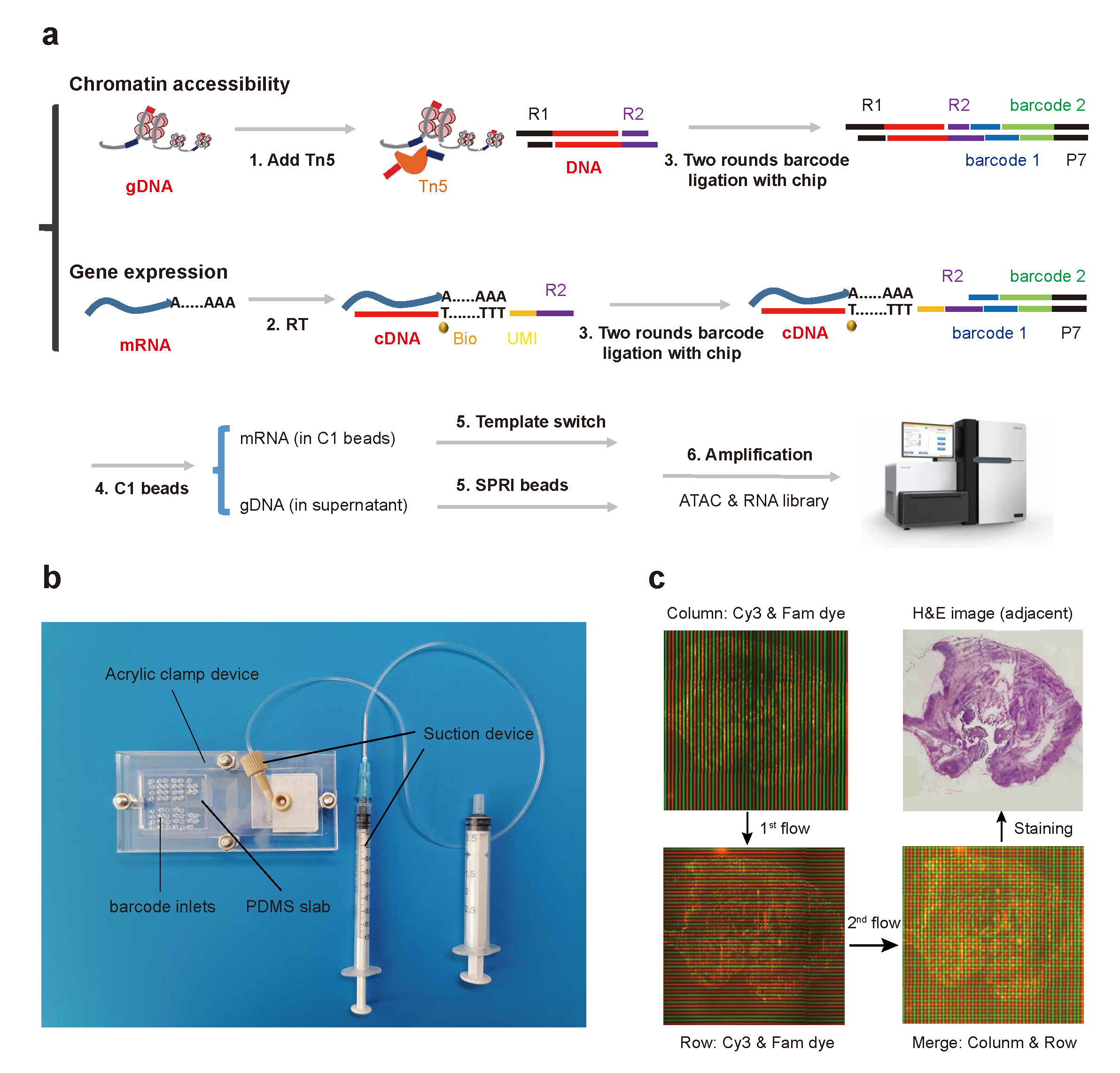
The first high-throughput spatial co-omics method, named as microfluidic indexing based spatial ATAC and RNA sequencing (MISAR-seq).
1. Download and install cellranger-arc software from 10x genomics and replace their default barcode file:
a. Enter cellranger-arc-2.0.1/lib/python/cellranger/barcodes/ and cellranger-arc-2.0.1/lib/python/atac/barcodes/ directory;
b. Replace default barcode file "737K-arc-v1.txt.gz" with your new custom barcode (in Barcode, also named as "737K-arc-v1.txt.gz").
bash MISAR-seq.sh
3. After fill tissue region with white and non-tissue with black in photoshop or other image processing software manually:
python Grid_filter.py
Rscript MISAR-seq.R
PS: New version in ZENODO: https://doi.org/10.5281/zenodo.7480069