Collections of examples of phylogenetic analysis with TreeTime
There are three different ways of using TreeTime:
- as a python module as part of python scripts and programs
- via a set of command-line tools
- via the webserver at treetime.ch.
The documentation of TreeTime is now hosted on readthedocs:
The general pattern to use TreeTime to infer timetrees via the command line is
treetime --tree mytree.nwk --aln mysequences.fasta --dates metadata.csvIn addition, there are number of subcommands for related tasks. Please see the following pages that explain usage for different cases:
- timetree estimation
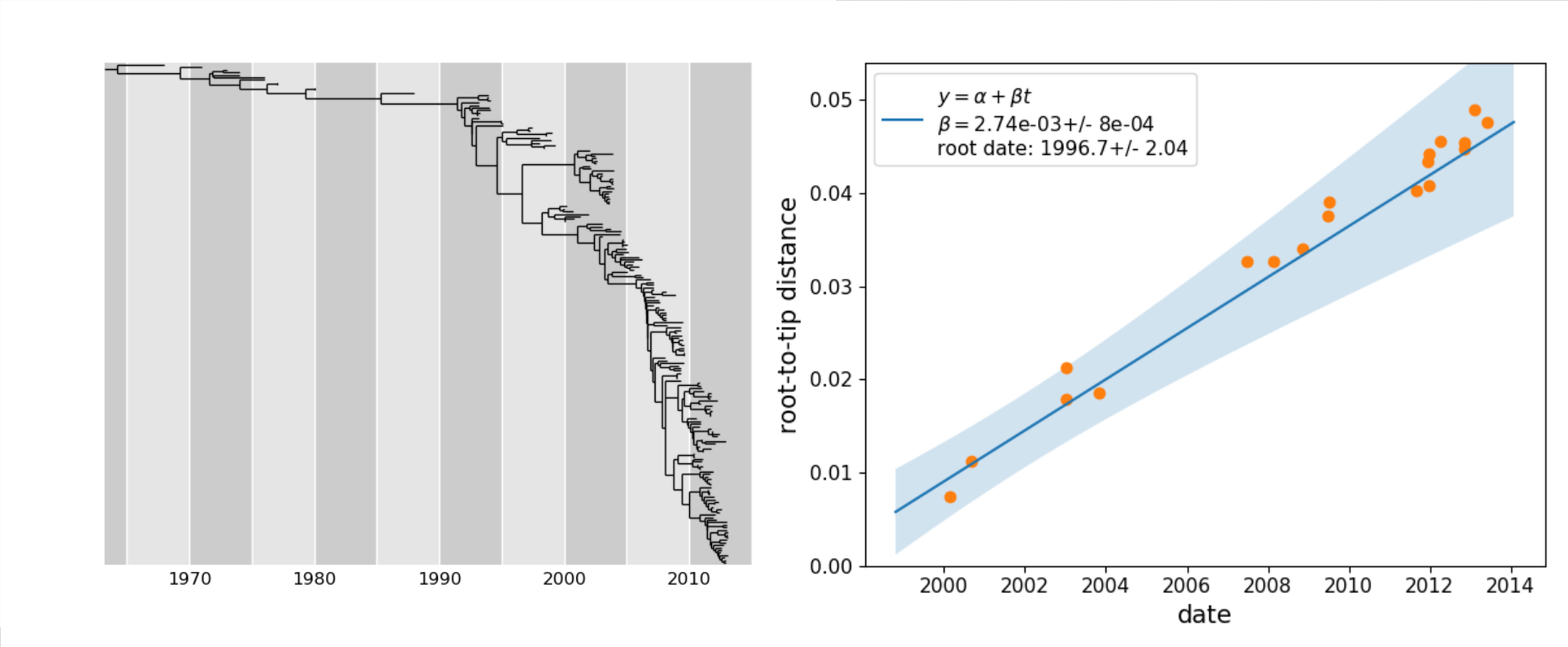
- estimation of substitution rates
- ancestral sequence reconstruction
- homoplasy analysis
- discrete traits and mugration models
A minimal script to run TreeTime would look something like this
from treetime import TreeTime
from treetime.utils import parse_dates
dates=parse_dates('metadata.csv')
tt=TreeTime(tree='mytree.nwk', aln='mysequences.fasta', dates=dates)
tt.run(root='best')More detailed examples can be found in the following set of scripts: