Hello,
I noticed an error in the WeisfeilerLehmanHashing class from utils.treefeatures.py which is used by Graph2Vec and Gl2vec for building the list of nodes rooted subtrees of the each graph up to the number of WL iterations (argument: wl_iterations).
I implemented a correction of WeisfeilerLehmanHashing and show you hereafter a comparison to demonstrate the problem.
reproducible code
It
(i) generates a simple chain graph
(ii) Run the WL algorithm implemented in your original code and print the WL features hash codes outputs.
(iii) Run the corrected algorithm and print the WL features hash codes outputs.
(iv) Print the WL nodes rooted trees associated with the WL features hash codes (they are in the same order).
import numpy as np
import networkx as nx
from karateclub.utils.treefeatures import WeisfeilerLehmanHashing
import hashlib
# We build a directed attributed graph (4 nodes chain)
A = np.matrix([[0,1,0,0],[0,0,1,0],[0,0,0,1],[0,0,0,0]])
G = nx.DiGraph(incoming_graph_data= A )
nx.set_node_attributes(G, {0:1,1:2,2:3,3:4}, name= 'feature')
print('Run original version')
WL = WeisfeilerLehmanHashing(graph= G, wl_iterations= 3, attributed=True)
print('Number of final WL features')
print(len(WL.get_graph_features()))
print('Final WL features')
print(WL.get_graph_features())
class CorrectedWeisfeilerLehmanHashing(object):
"""
Weisfeiler-Lehman feature extractor class.
Args:
graph (NetworkX graph): NetworkX graph for which we do WL hashing.
features (dict of strings): Feature hash map.
iterations (int): Number of WL iterations.
"""
def __init__(self, graph, wl_iterations, attributed):
"""
Initialization method which also executes feature extraction.
"""
self.wl_iterations = wl_iterations
self.graph = graph
self.attributed = attributed
#____ ADDED (For visualising trees only)
self.subtrees = []
self.all_subtrees = {}
#_______
self._set_features()
self._do_recursions()
def _set_features(self):
"""
Creating the features.
"""
if self.attributed:
self.features = nx.get_node_attributes(self.graph, 'feature')
else:
self.features = {node: self.graph.degree(node) for node in self.graph.nodes()}
#____ ADDED
self.extracted_features = {k: [str(v)] for k, v in self.features.items()}
#(For visualising trees only)
if self.attributed:
self.subtrees = nx.get_node_attributes(self.graph, 'feature')
else:
self.subtrees = {node: self.graph.degree(node) for node in self.graph.nodes()}
self.all_subtrees = {k:[v] for k,v in self.subtrees.items()}
#____
def _do_a_recursion(self):
"""
The method does a single WL recursion.
Return types:
* **new_features** *(dict of strings)* - The hash table with extracted WL features.
"""
#____ DELETED
#self.extracted_features = {k: [str(v)] for k, v in self.features.items()}
#____
#____ ADDED (For visualising trees only)
new_trees = {}
#_______
new_features = {}
for node in self.graph.nodes():
nebs = self.graph.neighbors(node)
degs = [self.features[neb] for neb in nebs]
features = [str(self.features[node])]+sorted([str(deg) for deg in degs])
features = "_".join(features)
hash_object = hashlib.md5(features.encode())
hashing = hash_object.hexdigest()
new_features[node] = hashing
#____ ADDED (For visualising trees only)
nebs = self.graph.neighbors(node)
neigbor_trees = [self.subtrees[neb] for neb in nebs]
ordered_neigbor_trees = [str(self.subtrees[node])]+sorted([str(tree) for tree in neigbor_trees])
new_node_rooted_tree = "("+"_".join(ordered_neigbor_trees)+")"
new_trees[node] = new_node_rooted_tree
#_______
self.extracted_features = {k: self.extracted_features[k] + [v] for k, v in new_features.items()}
#____ ADDED (For visualising trees only)
self.all_subtrees = {k : self.all_subtrees[k] + [v] for k,v in new_trees.items()}
self.subtrees = new_trees
#____
#____ ADDED
# we remove the initial non encoded feature for each node if still there
for k,v in new_features.items():
if len(self.extracted_features[k][0])!=32:
del self.extracted_features[k][0:1]
#____
return new_features
def _do_recursions(self):
"""
The method does a series of WL recursions.
"""
for _ in range(self.wl_iterations):
self.features = self._do_a_recursion()
#____ ADDED (For visualising trees only)
for k,v in self.all_subtrees.items():
del self.all_subtrees[k][0:1]
#____
def get_node_features(self):
"""
Return the node level features.
"""
return self.extracted_features
def get_graph_features(self):
"""
Return the graph level features.
"""
return [feature for node, features in self.extracted_features.items() for feature in features]
#____ ADDED (For visualising trees only)
def get_subtrees(self):
"""
Return the nodes rooted subtrees of all WL iterations
"""
return self.all_subtrees
print('Run corrected version')
WL = CorrectedWeisfeilerLehmanHashing(graph= G, wl_iterations= 3, attributed=True)
print('Number of final WL features')
print(len(WL.get_graph_features()))
print('Final WL features')
print(WL.get_graph_features())
print('Corresponding nodes rooted subtrees')
print(WL.all_subtrees)Here is the output of this script:
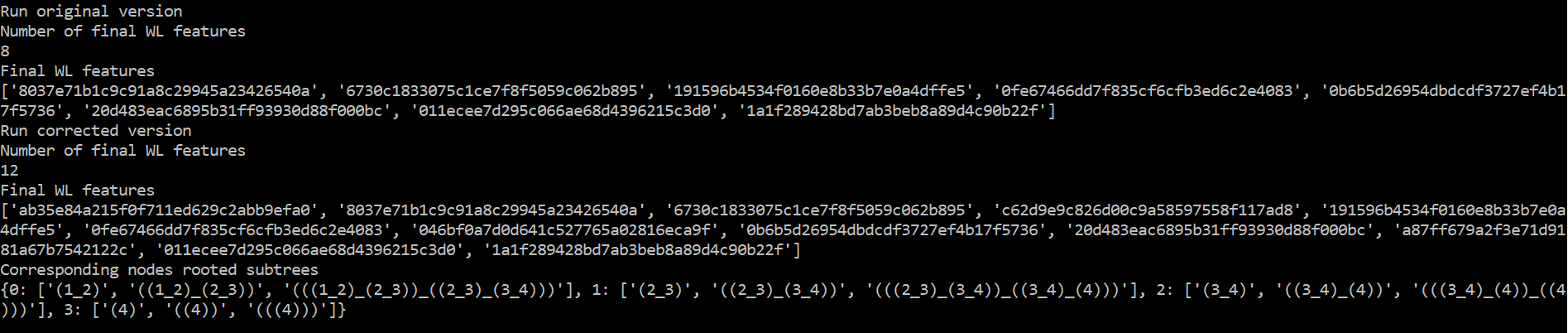
Explanation
Indeed, you can see first that the original code only produces 8 (4x2) WL features hash codes instead of 12 (4x3) in the corrected version. The algorithm must produce one WL feature per node and iteration.
You can also see that 8 codes among the corrected output match the non-corrected output. Now looking at the output of WL.all_subtrees, whose order match the features in WL.get_graph_features(), you can see that the codes that were lacking in the non-corrected version correspond to the first WL iteration.
Summary
The problem of the original code is that it only keep the WL features of the 2 last iterations. I hope that this is clear enough. Of course you can use directly the corrected version. I kept track of the changes and highlighted those that I only use for the trees strings construction.
Could you please apply the changes to the package? I'm using it for research and I think it would be much easier for everyone that I directly refer to your package rather than adding a corrected code in a personal repo.
Anyway, thank you for this useful package =)



