Paul A. Gagniuc's Projects
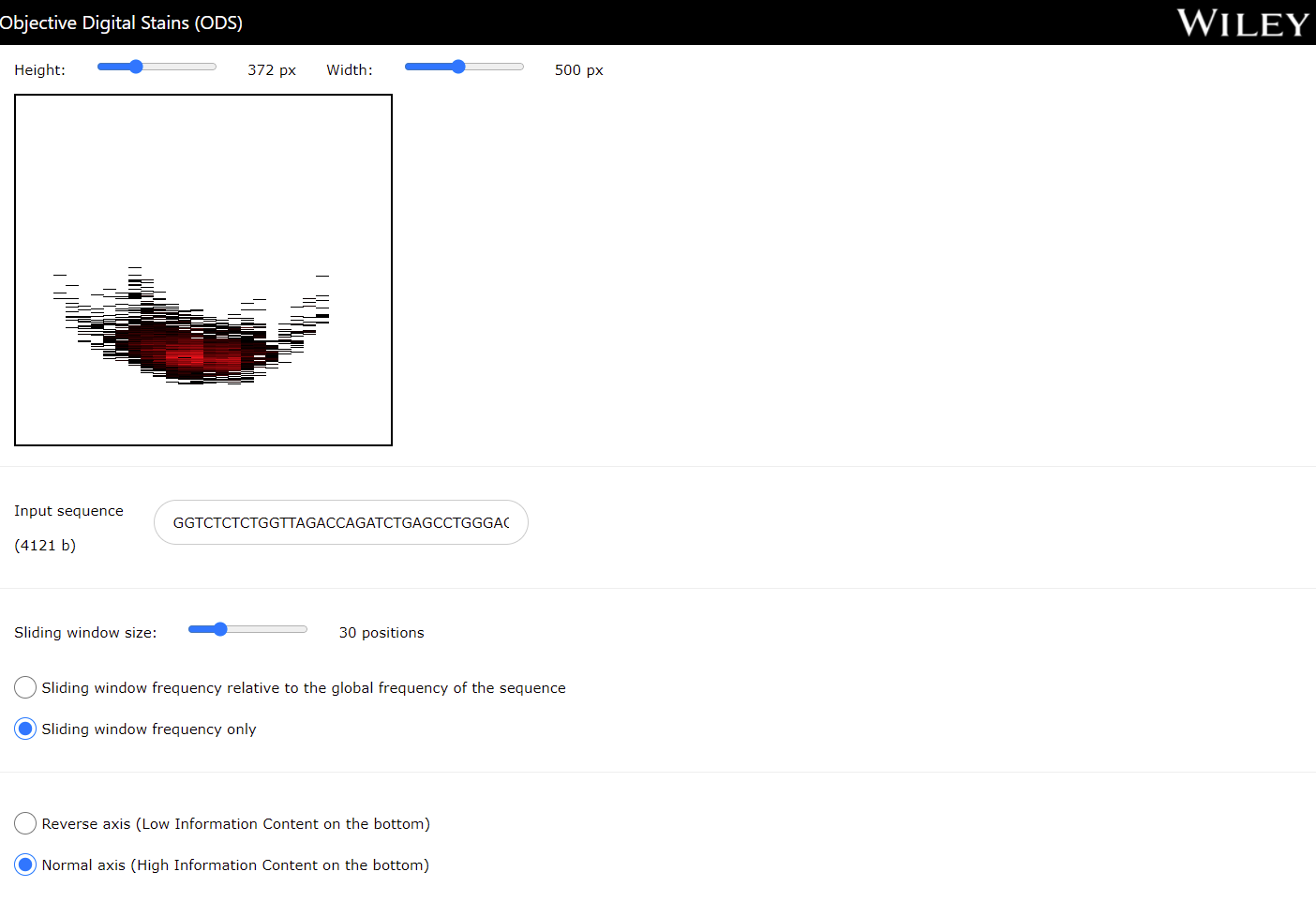
A 2D Objective Digital Stain is able to show the information structure of a DNA or RNA sequence in a graphical manner. In this case, the ODS is computed using the local frequency of the symbols from a sliding window.
A 3D Objective Digital Stain is able to show the information structure of a DNA or RNA sequence in a graphical manner. In this case, the ODS is computed using the local frequency of the symbols from a sliding window. In the 3D version, the overlapping values (similar sliding windows) are represented by a gradient from black to red.
A 3D Objective Digital Stain is able to show the information structure of a DNA or RNA sequence in a graphical manner. In this case, the ODS uses the global frequency of symbols (A, T/U, C, G) from the input sequence to calculate the local frequency of these symbols from a sliding window.
This is a scanner designed to recognise DNA motifs within a long stretch of DNA. It uses two models for discrimination, one model representing the target and the second model representing the background.
This is a high discrimination scanner designed to recognise DNA motifs within a long stretch of DNA. Most importantly, this implementation shows how to implement a variable sensitivity for detection, by modifying the pseudocount values.
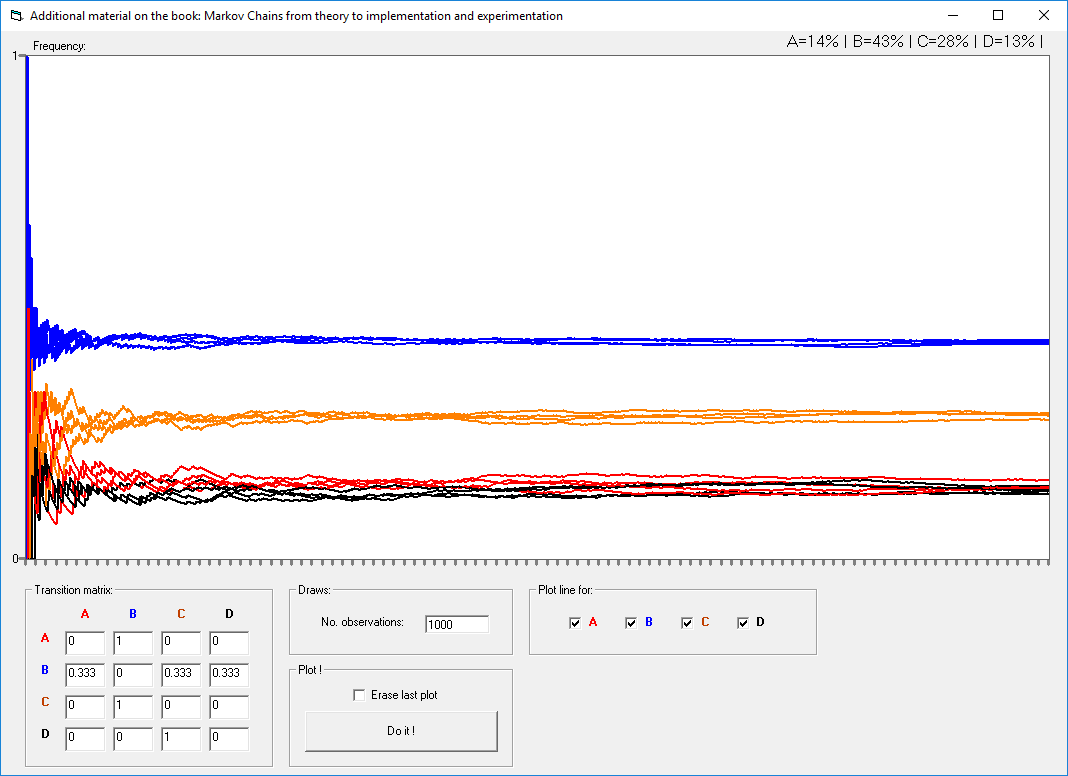
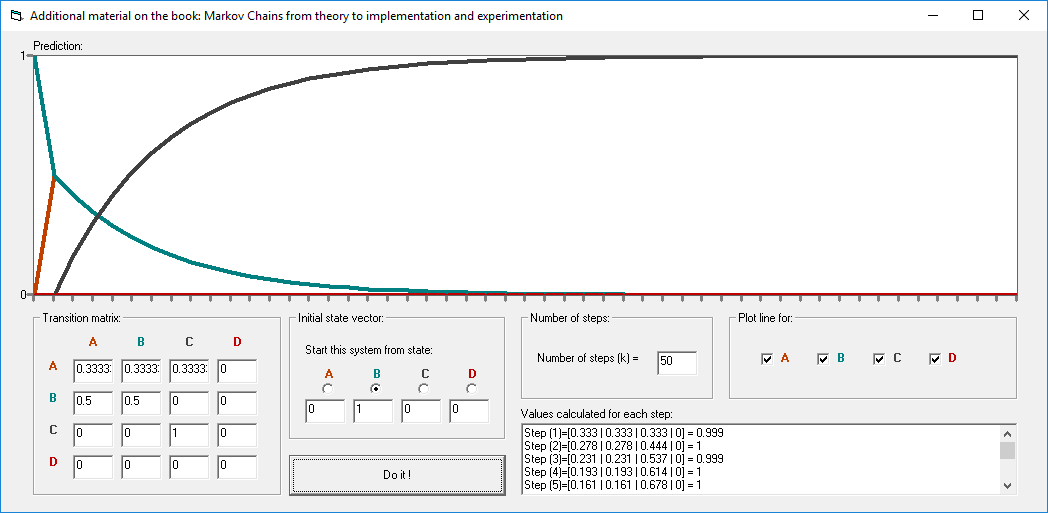
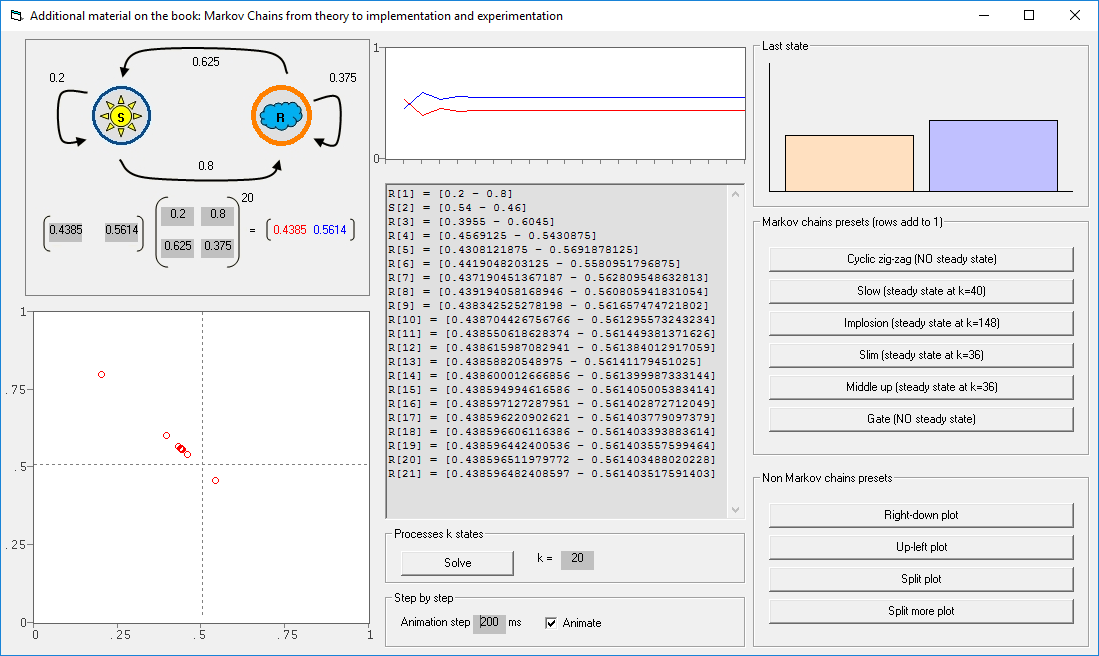
This repository includes the ".bas" files for Markov Chains that accompany the book entitled: Markov Chains: From Theory to Implementation and Experimentation. These ".bas" implementations can be used in various VBA Excel applications. This repository also includes an EXCEL file that supports VBA.
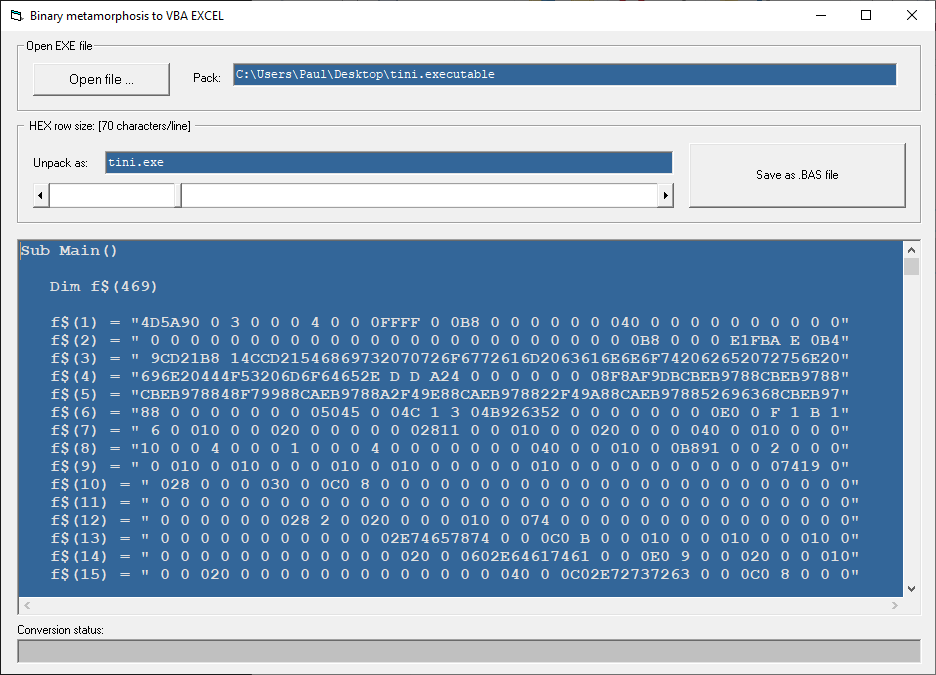
This application converts any executable file to VBA source code that can be included as a '.bas' module in an EXCEL file. Once inserted into the EXCEL file, the VBA code can be used to completely restore the executable file to disk in the same directory as the EXCEL file.
The VB6 applications shown here use the hexadecimal system to encode the binary content of an executable file. The point here is that one may compile an executable file that contains another executable file inside. Once the new executable file is executed, it is able to write the embedded executable file on disk as an independent executable file.
These Bioinformatics HTML5/JS files accompany the book entitled: Algorithms in Bioinformatics: Theory and Implementation, and they are compatible with all internet browsers. These algorithms include more than 120 open-source implementations.
These Bioinformatics HTML5/JS files accompany the book entitled: Algorithms in Bioinformatics: Theory and Implementation, and they are compatible with all internet browsers. These algorithms include more than 120 open-source implementations that describe many known or novel algorithms in Bioinformatics.
These All-in-one Bioinformatics HTML5/JS files accompany the book entitled: Algorithms in Bioinformatics: Theory and Implementation, and they are compatible with all internet browsers.
Chaos & noise is an implementation that suggests by experiment that the universe is deterministic. Initially, the experiment was used to show why there are limits in predicting the behavior of physical systems. The subject is openly described in the book entitled Algorithms in Bioinformatics: Theory and Implementation.
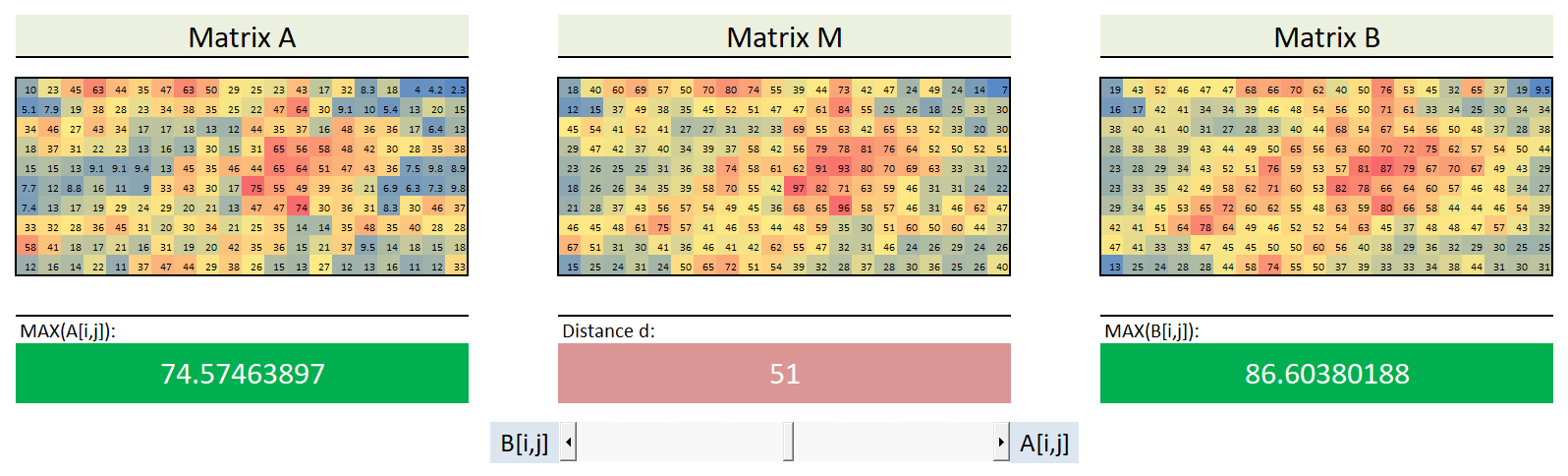
This is an implementation designed in eight different programming / scripting languages, namely C#, Python, VB6, Javascript, Perl, Ruby, Java and PHP. Each implementation is able to mix two signals/vectors (A and B) in arbitrary proportions.
Diabetes prediction V1.0 uses the Markov Chains method. First, this VB6 application converts a sequence of numbers into states. The states are arranged in a transition matrix and the transition probabilities are calculated for each element. Next, the transition matrix is further used for a prediction in a Markov chain.
Diabetes prediction V2.0 -This VB6 application takes glycemic values and tries to predict the future state of the patient. First, it converts a sequence of numbers into states. The states are arranged in a transition matrix and the transition probabilities are calculated for each element. The transition matrix is further used for a predictions.
These implementations make use of an algorithm called DPD (Discrete Probability Detector), that transforms any sequence of symbols into a transition matrix. DPD is able to detect the number of states from the sequence and calculate the transition probabilities between these states. This version of DPD is made in JavaScript, VBS and VB6.
Discrete Probability Detector (DPD) is an algorithm that transforms any sequence of symbols into a transition matrix. It is able to detect the number of states from the sequence and calculate the transition probabilities between these states. This version of DPD is made in Visual Basic 6.0.
Discrete Probability Detector (DPD) application uses an algorithm that transforms any sequence of symbols into a transition matrix. It is able to detect the number of states from the sequence and calculate the transition probabilities between these states. This version of DPD is made in JavaScript.
This JavaScript implementation detects the areas where two DNA sequences are complementary to each other. All symbols from UTF-8 are accepted by this algorithm.
This implementation is an alternative that provides full control over how the graphics of a Sequence Logo should look like, and is an alternative to an application called WebLogo. All the inner workings of this open source application are written in native javascript. The application is independent of the internet once it is saved as a html file.
The Dynamic Block Allocation algorithm (DBA) represents a flexible method for partitioning string sequences into data blocks taking into account different rules imposed by a function. Two versions of this algorithm are presented, namely DBFA (Double Brute Force Algorithm) and MBFA (Multi Brute Force Algorithm).
This application calculates the entropy of a string. The focus of this implementation is represented by a specialized function called "entropy" which receives a text sequence as a parameter and returns a value that represents the entropy. Entropy is a measure of the uncertainty in a random variable.
Entropy is a measure of the uncertainty in a random variable. This application calculates the entropy of text. The current example calculates the entropy of sequence "TTTAAGCC". In the context of information theory the term "Entropy" refers to the Shannon entropy.
Entropy vs self sequence alignment is a Javascript implementation of a scanner that makes a comparison between two methods, namely between Shanon entropy (Information entropy) and self-sequence alignment (Information content). Information entropy (IE) and Information content (IC) are two methods that quantitatively measure information.
Config files for my GitHub profile.
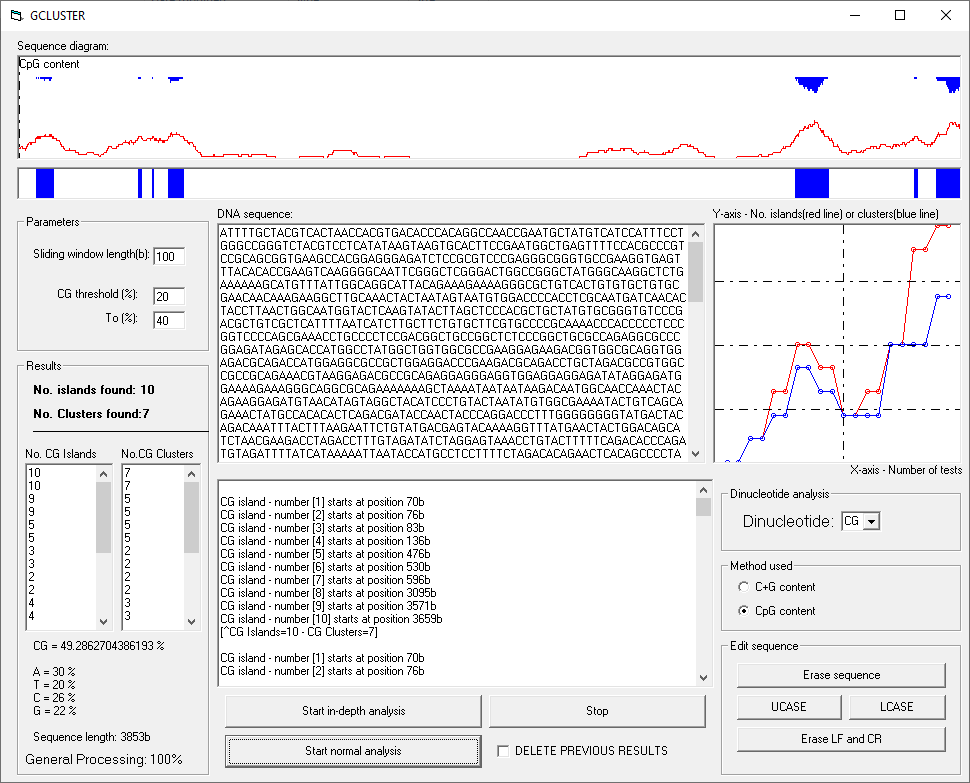
GCluster is an experimental detector that uses a dynamic method named "in-depth analysis" to detect and interpret CpG islands, CpG clusters and other dinucleotide structures. In-depth analysis is made through repeated tests with different dinucleotide thresholds.
Genomin is an implementation for large-scale genomic analysis. It is made in Visual Basic 6.0 (VB6). It uses the seek method to generate buffers from large FASTA files (over 8 Gb).
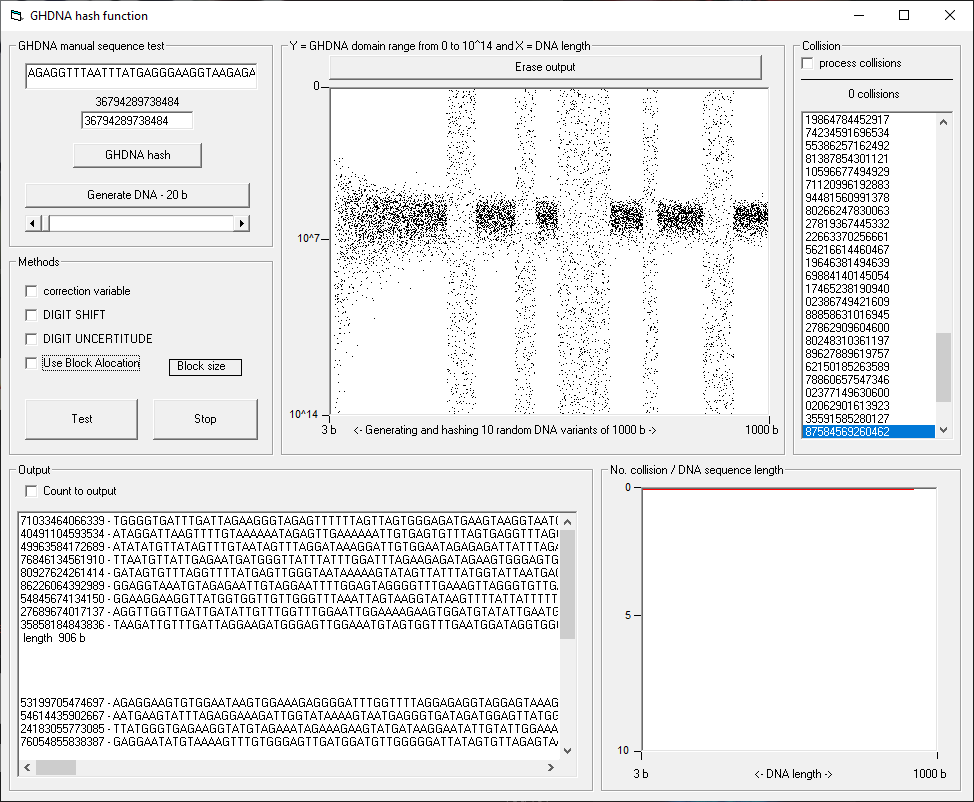
GHDNA is an experimental hash function that takes a DNA sequence as an input and provides a unique signature in the output. The signature provided by the function is a constant length sequence of digits.




.png?raw=true)
.png?raw=true)
.png?raw=true)








.png?raw=true)
%20with%20variable%20sensitivity.png?raw=true)







.png?raw=true)
.png?raw=true)
.png?raw=true)
.png?raw=true)
.png?raw=true)
.png?raw=true)




.png?raw=true)
.png?raw=true)
.png?raw=true)
.png?raw=true)