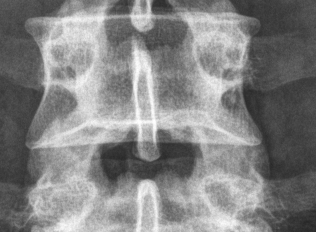
Simple open-source multi-platform DICOM browser/viewer in Python and Qt.
- Easy (and fast) viewing of all images in a directory
- Zooming (1:N and N:1)
- Mouse control of window center and width (as in Aeskulap Viewer)
- Proper measurement of Hounsfield units and position by mouse
- PNG image export
- Better zooming
- Better MRI images support
- RT dose images support
- View in different planes (rectangular + others)
- Coordinate mapping (using translation and rotation matrix)
- Integration of anonymization features (see https://github.com/janpipek/anonymize_dicom )
- Information from the DICOM file in user-friendly display
- Python 3 / Python 2.7 (not tested)
- qtpy (and therefore PyQt4 / PyQt5 / PySide - not automatically installed by pip!)
- pydicom (0.9.9, maybe also 1.0)
Tested on Linux and Windows.
The easiest way is pip install pydiq.
Currently, the viewer supports only Computed Radiography (CR), Computed Tomography (CT) and Magnetic Resonance Imaging (MRI) images with normal orientation (x, y, z) in one-slice-per-file format.